Tree Kernels In SVM-lighT
by
Alessandro Moschitti
The
latest version of the software is SVM-lighT-TK 1.5
Syntactic
parsers
are among of the most useful tools for Natural Language
Processing (NLP) applications.
However, how to exploit the syntactic parse tree
information in NLP tasks is considered an open problem.
For example, the learning models for automatic Word
Sense Disambiguation or Coreference Resolution would benefit
from syntactic tree features but their design and selection is
not an easy task.
Convolution
kernels
(see Kernel philosophy in NLP)
are an alternative to the explicit feature design. They
measure similarity between two syntactic trees in terms of
their sub-structures (e.g. [Collins and Duffy, 2002]). These
approaches have given optimal results [Moschitti, 2004] when
introducing syntactic information in the task of Predicate
Argument Classification.
Suppose
we want to measure the similarity between the parse trees of
two noun phrases: "a dog" and "a cat". The idea of behind
syntactic tree kernels is graphically described by the
following figure:
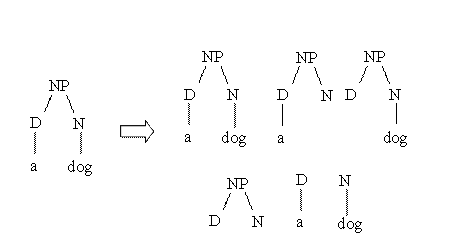
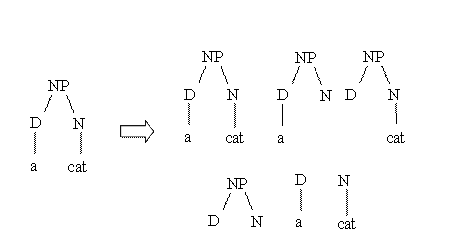
As
3 structures (out of 5) are completely identical the
similarity is equal to 3.
This
kind of similarity has been shown useful to improve the
ranking of the m best
syntactic parse trees [Collins and Duffy, 2002]. Other
interesting applications concern the classification of
predicate argument structures annotated in PropBank
and Question Classification [Zhang
and Lee, 2003;
Moschitti, ECML 2006]. To describe the linguistic phenomenon
of the former task, i.e. the syntactic/semantic relation
between a predicate and the semantic roles of its arguments,
we need to extract features from some subparts of a syntactic
tree. For example given the following sentence:
Paul
gives a talk in Rome
The
Predicate
argument annotation in PropBank is:
[
Arg0 Paul] [
Predicate
gives] [ Arg1
a talk] [ ArgM
in
The syntactic tree may be:
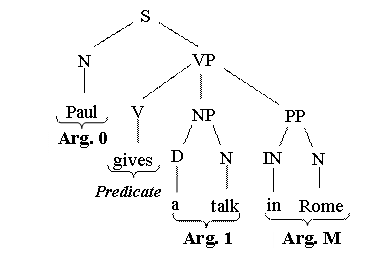
To
select
syntactic information related to a particular argument type we
could use the corresponding subtree, i.e.:
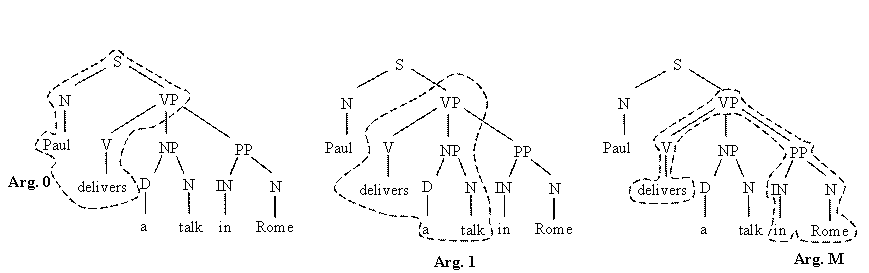
Tree-Kernels measure the similarity between arguments whereas the kernel-based algorithms (e.g. SVMs) automatically select the substructures that better describe the argument type.
How to use SVM-lighT-TK
The tree kernel has been encoded inside the well known SVM-light software written by Thorsten Joachims (www.joachims.org). The input format and the new options are compatible with those of the original SVM-light 5.01. Moreover, combination models between tree kernels and feature vectors are available.
Software Features
-
Fast Kernel Computation [Moschitti, EACL 2006]
(already available since the previous version).
-
Vector sets, multiple feature vectors over multiple feature spaces can
be specified in the input. This allows us to use different
kernels with different feature subsets. Given two objects, O1
and O2,
described by two sets of feature vectors, ![]() and
and
![]() ,
several kernels can be
defined:
,
several kernels can be
defined:
![]()
-
Tree forests, a set of trees over multiple feature spaces can be
specified in the input. This allows us to use a set of
different structured features, e.g. it is possible to extract
different portions from a parse tree and combine the different
contributions. This limits the sparseness of the kernels
applied to the whole tree. Given two objects, O1
and O2,
described by two sets of trees, ![]() and
and
![]() ,
several kernels can be
defined on:
,
several kernels can be
defined on:
![]()
-
Two types of tree kernels:
1.
SubSet
Tree
kernel (SST) [Collins and Duffy, 2002; Moschitti,
EACL 2006]
2.
Subtree
kernel
(ST) [Vishwanathan and Smola, 2001; Moschitti,
EACL
2006]
-
Embedded combinations of Trees and vectors:
1.
sequential
summation,
the kernels between corresponding pairs of trees and/or
vectors in the input sequence are summed together. The t
parameter rules the contributions of tree kernels Kt
with respect to the feature vector kernel kb.
Each of the two types of kernels can or cannot be normalized
according to a command line parameter. More formally:
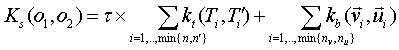 ,
,
Kt can be either the SST or the ST kernel whereas kb can be one of the traditional kernels on feature vectors, e.g. gaussian or polynomial kernel.
2.
all vs all summation, each tree and vector of the first object are evaluated
against each tree and vector of the second object:
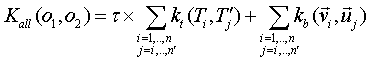
Data Format
The data format is the same as SVM-Light-TK-1.2. I improved
the generality of the tree please read the explanation of the
software below.
The major difference is the use of more general trees and the
possibility to execute the Partial Tree Kernel, (PTK)
[Moschitti, 2006].
You
can use any tree for example the simple tree below
A
/ | \
a b c
can
be represented with the following syntax:
(A (a)(b)(c))
Pay attention!! you cannot write such tree as (A a b c).
More in detail the input takes sets of objects:
<line> ::= <target><blank><set-of-vectors>
|
<target><blank><set-of-trees> |
<target><blank><trees-and-vectors>
<set-of-vectors>
::= <vector>
|<vector><blank><begin-vector><blank><vector><blank>..<end-vector>
<set-of-trees>
::= <begin-tree><blank><tree><blank>..<begin-tree><blank><tree><blank><end-tree>
<trees-and-vectors>::=
<set-of-trees><blank><set-of-vectors>
<vector> ::=
<feature>:<value><blank><feature>:<value><blank>...<blank><feature>:<value>
|
<blank>
<target>
::= +1 | -1 | 0 | <float>
<feature>
::= <integer> | "qid"
<value>
::= <float>
<begin-tree>
::="|BT|"
<end-tree>
::="|ET|"
<begin-vector>::="|BV|"
<end-vector>
::="|EV|"
<tree> ::= <full-tree> | <blank>
<full-tree>
::=
(<root><blank><full-tree>..<full-tree>)
| (<root><blank><leaf>)
<leaf> ::= <string>
<root> ::= <string>
<blank> ::= " " (i.e. one space)
where <string> does not contain space, left and right parentheses, i.e. "(" and ")", whereas <tree> defines the usual Penn Treebank format. Note that (a) two begin trees, i.e. ".. |BT| |BT| .." encode the empty tree (useful as placeholder), (b) two begin vectors, i.e. ".. |BV| |BV| .." encode the empty vector and (c) the sequence ".. |ET| |BV| .." is used to specify that the first vector (after trees) is empty.
For example, if we liked to experiment with different trees for question classification, given the question "What does S.O.S stand for?", we may use the following forest:
1 |BT| (SBARQ (WHNP (WP What))(SQ (AUX does)(NP (NNP S.O.S.))(VP (VB
stand)(PP (IN for))))(. ?)) |BT|
(BOW (What
*)(does *)(S.O.S. *)(stand *)(for *)(? *)) |BT|
(BOP (WP
*)(AUX *)(NNP *)(VB *)(IN *)(. *)) |BT|
(PAS (ARG0 (R-A1 (What *)))(ARG1 (A1 (S.O.S. NNP)))(ARG2 (rel
stand))) |ET|
where the four different trees are: the question parse tree, the BOW tree (used to simulate the bag-of-words), the BOP tree (used to simulate the bag-of-POS-tags) and the predicate argument tree (i.e. the PAS defined in [Moschitti et al., ECML-MLG 2006])
We
can add trees with different feature vectors. For example
suppose we want to implement a re-ranker based on tree kernel
and flat features. We need to compare pairs of instances. The
following line contains a pair of PASs and a pair of feature
vectors that correspond to two target predicate argument
structures. These are needed for learning of a re-ranker for
Semantic Role Labeling systems [Moschitti et al., CoNLL 2006].
-1 |BT| (TREE
(ARG0 (A1 NP))(ARG1 (AM-NEG RB))(ARG2 (rel fall))(ARG3 (AM-TMP
NNP))(ARG4 (AM-TMP SBAR))(ARG5 null)(ARG6 null)) |BT|
(TREE (ARG0 (A1 NP))(ARG1 (AM-NEG RB))(ARG2 (rel fall))(ARG3
(A4 RP))(ARG4 (AM-TMP NNP))(ARG5 (AM-TMP SBAR))(ARG6 null)) |ET| 1:1 21:2.742439465642236E-4 23:1 30:1 36:1 39:1 41:1 46:1 49:1
66:1 152:1 274:1 333:1 |BV|
2:1 21:1.4421347148614654E-4 23:1 31:1 36:1 39:1 41:1 46:1
49:1 52:1 66:1 152:1 246:1 333:1 392:1 |EV|
In
case we liked to use only feature vectors we could write them
as follows:
-1 1:1 21:2.742439465642236E-4 23:1 30:1 36:1 39:1 41:1 46:1
49:1 66:1 152:1 274:1 333:1 |BV|
2:1 21:1.4421347148614654E-4 23:1 31:1 36:1 39:1 41:1 46:1
49:1 52:1 66:1 152:1 246:1 333:1 392:1 |EV|
However,
the original SVM-light input format can be used without any
changes as the next two lines associated with two instances
illustrate:
-1 1:1 21:2.742439465642236E-4 23:1 30:1 36:1 39:1 41:1 46:1
49:1 66:1 152:1 274:1 333:1
+1 2:1 21:1.4421347148614654E-4 23:1 31:1 36:1 39:1 41:1
46:1 49:1 52:1 66:1 152:1 246:1 333:1 392:1
Important:
follow the syntax of trees defined by <tree> (e.g.
there
is no space between two parenthesis). Moreover,
the expected input is a parse-tree this means that a
pre-terminal (the lowest non-terminal in the tree) is always
followed by only one leaf.
Commands
The "svm_classify" and the "svm_learn" commands maintain the original svm-light format:
usage:
svm_learn
[options] example_file model_file
Arguments:
example_file-> file with training data
model_file ->
file to store the learned decision rules in
usage:
svm_classify
[options] example_file model_file
Arguments:
example_file-> file with testing data
model_file ->
file to retrieve the learned decision rules
The
new options are shown below (in blue colour):
Kernel
options:
-t int ->
type of kernel function:
0:
linear (default)
1:
polynomial (s a*b+c)^d
2:
radial basis function exp(-gamma ||a-b||^2)
3:
sigmoid tanh(s a*b + c)
4:
user defined kernel from kernel.h
5:
combination of forest and vector sets according to W, V, S, C
options
11:
re-ranking based on trees (each instance must have two trees)
12:
re-ranking based on vectors (each instance must have two
vectors)
13:
re-ranking based on both tree and vectors (each instance must
have two trees and two vectors)
-W
[S,A] -> a tree
kernel is applied to the sequence of trees of two input
forests and the results are summed;
-> with an "A", a tree kernel is applied to all tree
pairs from the two forests (default "S")
-V
[S,A] -> same as
before but sequences of vectors are used (default "S" and the
type of vector-based kernel is specified by the option -S)
-S
[0,4] -> kernel to be
used with vectors (default polynomial of degree 3, i.e. -S = 1
and -d = 3)
-C
[*,+,T,V] ->
combination operator between forests and vectors (default 'T')
-> "T" only the contribution from trees is used
-> "V" only the contribution from feature vectors is
used
-> "+" or "*" sum or multiplication of the
contributions from feature vectors and trees (default 'T')
-T
float ->
multiplicative constant for the contribution of tree kernels
when -C = "+", i.e. K = tree-forest-kernel*r + vector-kernel
(default 1)
-D [0,1] -> 0, SubTree kernel or 1, SubSet Tree kernels (default 1)
-F
[0,1,2,3,4,6]-> 0 = ST kernel, 1 = SST kernel, 2 = SST-bow,
3 = PT kernel (default 1), 6=STRING kernel
-M
float -> Mu decay factor for PT kernel (default 0.4)
-L
float -> decay factor
in tree kernels (default 0.4)
-N [0,3] -> 0 = no normalization, 1 = tree normalization, 2 = vector normalization and, 3 = normalization of both trees and vectors. The normalization is applied to each individual tree or vector (default 3).
-u
string -> parameter
of user defined kernel
-d int -> parameter d in
polynomial kernel
-g
float -> parameter
gamma in rbf kernel
-s
float -> parameter s
in sigmoid/poly kernel
-r
float -> parameter c
in sigmoid/poly kernel
To obtain the sequential summation Ks (of tree and vector kernels) previously defined, we can set the option "-t 5 -T 1 -W S -V S -C +". Considering the default values, this is equivalent to use "-t 5 -C +".
-F
decides the type of tree kernel
-1
// Moschtti et al. 2007
0
//
VISHTANAM and SMOLA 2002
1 // COLLINS and DUFFY 2002
2
// ZHANG, 2003
3 // Moschitti, ECML 2006
4
// something ibrid
6 // STRING_KERNEL, Taylor and Cristianini book 2004, just put a fake root and only one level of children, i.e., the elements of your sequence.
Example of New Options
./svm_learn
-t
5 example_file model_file /* the subset-tree kernel alone is
used, if the forest contains only a tree, the classic tree
kernel is computed */
./svm_learn
-t
5 -C V example_file model_file /* the default polynomial
kernel is used on the pairs from vector sequences */
./svm_learn
-t
5 -C V -V A example_file model_file /* the default polynomial
kernel is used on the pairs from
vector
sequences.
The pairs are built by combining each element of the first
sequence with each element of the second sequence */
./svm_learn -t 5 -C + -S 1 -d 5 example_file model_file /* the sequential summation of trees, using SST kernel, is summed to the sequential summation of vectors, using a polynomial kernel with degree = 5. The contribution of tree kernels is multiplied by t (i.e. default 1) */
./svm_learn -t 5 -C + -D 0 -S 1 example_file model_file /* the sequential summation of trees, using the ST kernel (-D 0), is summed to the sequential summation of vectors, using polynomial kernel with degree = 5 */
./svm_learn -t 12 example_file model_file /* a re-ranker over a pair of trees and a pair of vectors is applied*/
./svm_learn
example_file
model_file /* original SVM-light linear kernel "-t 0". The
input can be provided in the new style or in the old SVM-light
format*/
Download SVM-LIGHT-TK 1.5
- Source code (It is available with the make files for Windows DevC++ and Linux gcc)
- Example data (it contains the PropBank Argument 0 as positive class and Argument 1 as negative class)
It is possible to design our own kernel by using weights for both trees and vectors and combining them in very different way as the following example illustrates:
- kernel.h
Download SVM-LIGHT-TK 1.2
This version does not contain PTK, it should not be faster than the new SVM-Light-TK1.5. Anyway if you do not need general trees and PTK, you may want to use this one.
- Source code (It is available with the make files for Windows DevC++ and Linux gcc)
Download SVM-LIGHT-TK
1.0
All the options of previous
SVM-LIGHT-TK version have been integrated in SVM-LIGHT-TK 1.2.
However, the use of forest and vector set make slightly slower the new version so if you just need a fast computation of one tree kernel.
Note that the data format is
different so please watch the page:
More
documentation examples are available at the link of my
Tutorial on kernels at
Tutorial: Kernel-based Learning to
Rank with Syntactic and Semantic Structures
ACL State-of-the-Art
Kernels for Natural Language Processing
Software
References
Please
refer
to the tree-kernel software as one of the following
[Moschitti, 2006], Alessandro Moschitti, Efficient Convolution Kernels for Dependency and Constituent Syntactic Trees. In Proceedings of the 17th European Conference on Machine Learning, Berlin, Germany, 2006.
[Moschitti, 2004], Alessandro Moschitti. A
study on Convolution Kernels for Shallow Semantic Parsing.
In proceedings of the 42-th Conference on Association for
Computational Linguistic (ACL-2004),
[Moschitti, 2006], Alessandro Moschitti,
Making tree kernels
practical for natural language learning. In
Proceedings of the Eleventh International Conference on
European Association for Computational Linguistics,
along
with
the SVM-light software as
[Joachims,
1999], Thorsten Joachims. Making large-scale SVM
learning practical. In B. Scholkopf, C. Burges, and A. Smola,
editors, Advances in Kernel Methods - Support Vector Learning,
1999.
General
References
[Collins and Duffy, 2002], Michael Collins and Nigel Duffy. New ranking algorithms for parsing and tagging: Kernels over discrete structures, and the voted perceptron. In ACL02, 2002.
[Moschitti, ECML 2006], Alessandro Moschitti,
Efficient Convolution
Kernels for Dependency and Constituent Syntactic Trees.
In Proceedings of the 17th European Conference on Machine
[Moschitti et al., ECML-MLG 2006], Alessandro Moschitti, Daniele Pighin, and Roberto Basili, Tree Kernel Engineering for Proposition Re-ranking, In Proceedings of Mining and Learning with Graphs (MLG 2006), Workshop held with ECML/PKDD 2006, Berlin, Germany, 2006.
[Moschitti et al., CoNLL
2006], Alessandro Moschitti, Daniele Pighin and Roberto
Basili. Semantic Role Labeling via
Tree Kernel joint inference.
In Proceedings of the 10th Conference on Computational Natural
Language
[Vishwanathan and Smola, 2002], S.V.N. Vishwanathan and A.J. Smola. Fast kernels on strings and trees. In Proceedings of Neural Information Processing Systems, 2002.
[Zhang and Lee, 2003], Zhang, D., Lee, W.S.: Question classification using support vector machines. In: Proceedings of SIGIR'03, 2003.
|
Maintained
by Alessandro Moschitti moschitti[at]dit.unitn.it |